Could not wait for the missing Elixir ML libraries to appear, so, I wrote one myself, taking https://github.com/sdwolfz/exlearn as a foundation.
The name is Matrex and it’s super-fast (compared to pure Elixir implementations) matrix manipulation lib.
Critical code is written in C using CBLAS subroutines and linked as Erlang NIFs.
It’s about 50-5000 times faster, than pure Elixir.
In the repo you will find MathLab fmincg() ported to Elixir with the help of the library
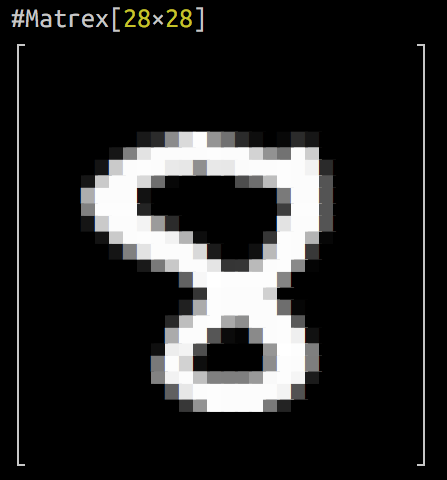
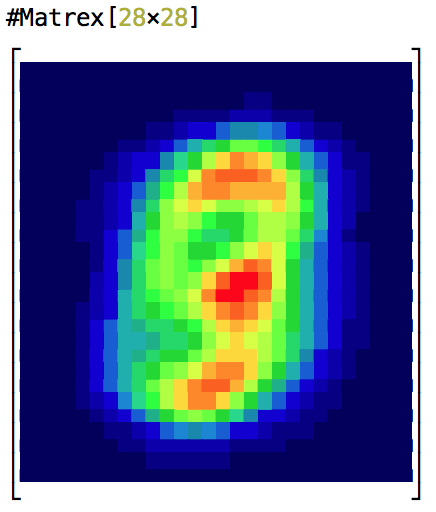
and logistic regression MNIST digits recognition exercise from Andrew Ng’s ML course implemented in Elixir (15 times faster, than Octave implentation).
It can be used like this:
y = Matrex.load("y.mtx")
j =
y
|> Matrex.dot_tn(Matrex.apply(h, :log), -1)
|> Matrex.substract(
Matrex.dot_tn(
Matrex.substract(1, y),
Matrex.apply(Matrex.substract(1, h), :log)
)
)
|> Matrex.scalar()
|> (fn
NaN -> NaN
x -> x / m + regularization
end).()
Or like this:
import Matrex.Operators
h = sigmoid(x * theta)
l = ones(size(theta)) |> set(1, 1, 0.0)
j = (-t(y) * log(h) - t(1 - y) * log(1 - h) + lambda / 2 * t(l) * pow2(theta)) / m
I’ve also created a Jupyter notebook with logistic regression algorithm in Elixir built with the help of this library.
Please, check Matrex on GitHub,
take a look at Matrex hex docs,
and tell me what you think of it.