Chapter 1 is read!
This was a great read. I don’t have any prior exposure to genetic algorithm, other than my university AI course that happened 20 years ago. So, this was quite a fun experience for me.
My initial reaction was to read the prose first and take a stab at the code later, but the way the example was arranged, I felt it would be more convenient if I experiment with each function and their result on a shell, and what better shell than LiveBook?
I experimented with each artifacts (i.e. population, evaluate, selection, crossover, mutation (when it appeared after the interval), algorithm). I even came up with an acronym (P.E.S.C.A), until M for mutation came along and mutated my acronym into an unpronounceable one. I keep thinking evaluate could evaluation to belong to same parts of speech class as all other functions (or selection → select, mutation → mutate etc). Or maybe that’s just me.
I really liked how, for the first time in my life, I understood what an AI book is trying to tell me in one run. The writing style is enjoyable and the introductory elements fit right in. The presentation, both in terms of english and code, are very easy to visualize. I have not seen any genetic algorithm code in other languages but I felt like both the recursion and piped workflow is perfect representation. I eagerly await reading next chapters to see if this can turn into a generic pattern for larger set of problems.
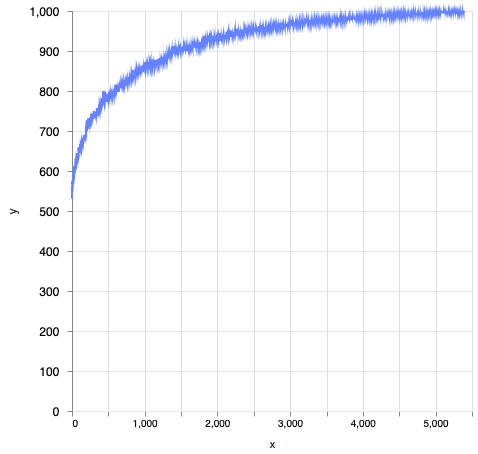
I have discussed the code running completely without mutate (or is it mutation?) at all attempts I made (contrary to what the book said), but one update is that, when I tried the same on 10, 100 on REPLIT (1 CPU, OTP 24 etc), then it kept dancing around 82, 87 for 200k iterations  … but that aside, running all the code is strongly recommended if you ask me, even better with LiveBook. It gives a “feel” for what the author is trying to explain.
… but that aside, running all the code is strongly recommended if you ask me, even better with LiveBook. It gives a “feel” for what the author is trying to explain.
That’s my brief summary of my experience with chapter 1. A bunch of things I would like to do (Please folks, tell me if this would be futile):
- Create a repository of livemds for the code
- Try using the
Nx data structures and functions to solve the same problems, who knows, if chapter permits, Explore?
- Produce
VegaLite charts.
- Benchmark things on different backends (I will admit, I don’t even know what backends will do for these problems)
Anyways, I will try to pick one or more of those and share it with y’all. That’ll be all for my chapter 1. I briefly read chapter 2 and already like the way it built upon Chapter 1. Thank you @AstonJ and @stevensonmt for doing this.
![]()
![]()